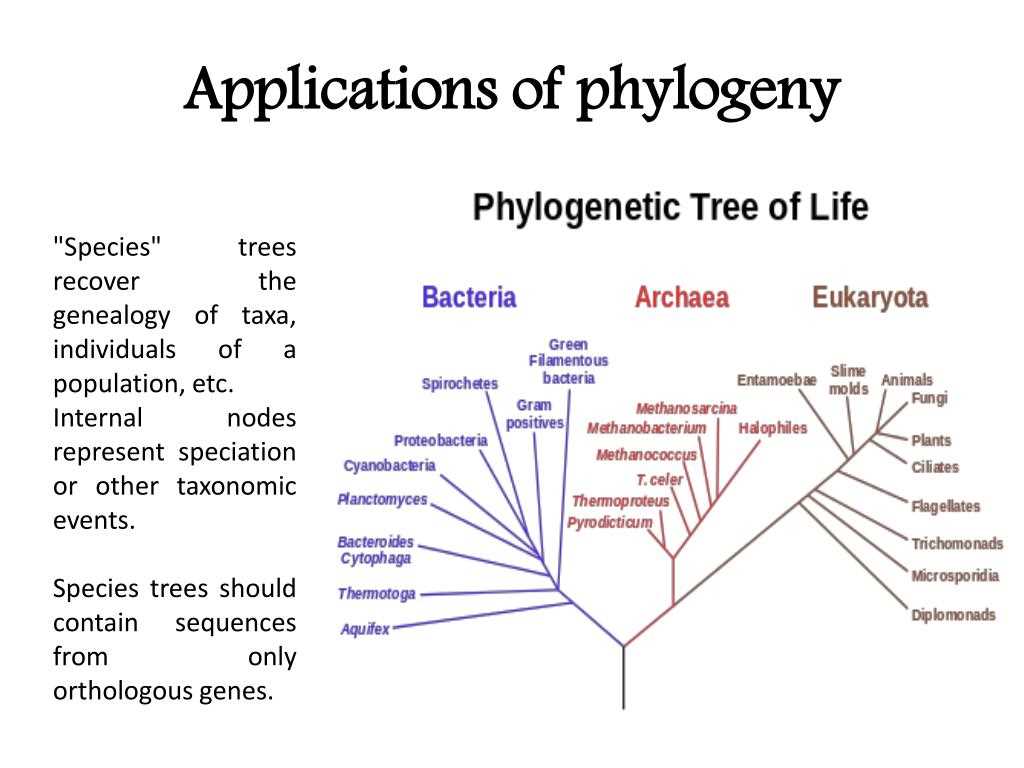
In the study of evolutionary biology, phylogenetic trees play a crucial role in understanding the relationships between different species. These trees depict the evolutionary history of organisms by analyzing their DNA sequences. Phylogenetic trees are constructed based on the idea that species that share more similar DNA sequences are more closely related.
To create phylogenetic trees, scientists first collect DNA samples from different species. They then isolate and sequence specific regions of the DNA, such as the mitochondrial DNA or certain genes. The sequences are then aligned and compared to identify similarities and differences.
Once the DNA sequences have been obtained and aligned, scientists use various computational tools and algorithms to construct the phylogenetic tree. One commonly used method is the Maximum Likelihood method, which calculates the probability of different evolutionary paths based on the observed DNA sequence data.
Creating phylogenetic trees from DNA sequences can provide insights into many aspects of evolution, such as the common ancestors of different species, the timing of evolutionary divergences, and the process of speciation. These trees can also be used to study the genetic relationships between different populations within a species and to track the spread of diseases.
In conclusion, creating phylogenetic trees from DNA sequences is a powerful tool in evolutionary biology. It allows scientists to unravel the intricate relationships between different species and gain a better understanding of the mechanisms driving evolution. By analyzing DNA sequences, scientists can reconstruct the evolutionary history of organisms and make important discoveries about the diversity of life on Earth.
Overview of Phylogenetic Trees
Phylogenetic trees are diagrams that represent the evolutionary relationships among different species or groups of organisms. These trees are constructed using genetic data, such as DNA sequences, and are based on the principle of common ancestry. Phylogenetic trees can provide valuable insights into the evolutionary history and relationships of organisms, and they are widely used in fields such as biology, ecology, and paleontology.
Key Concepts and Terminology
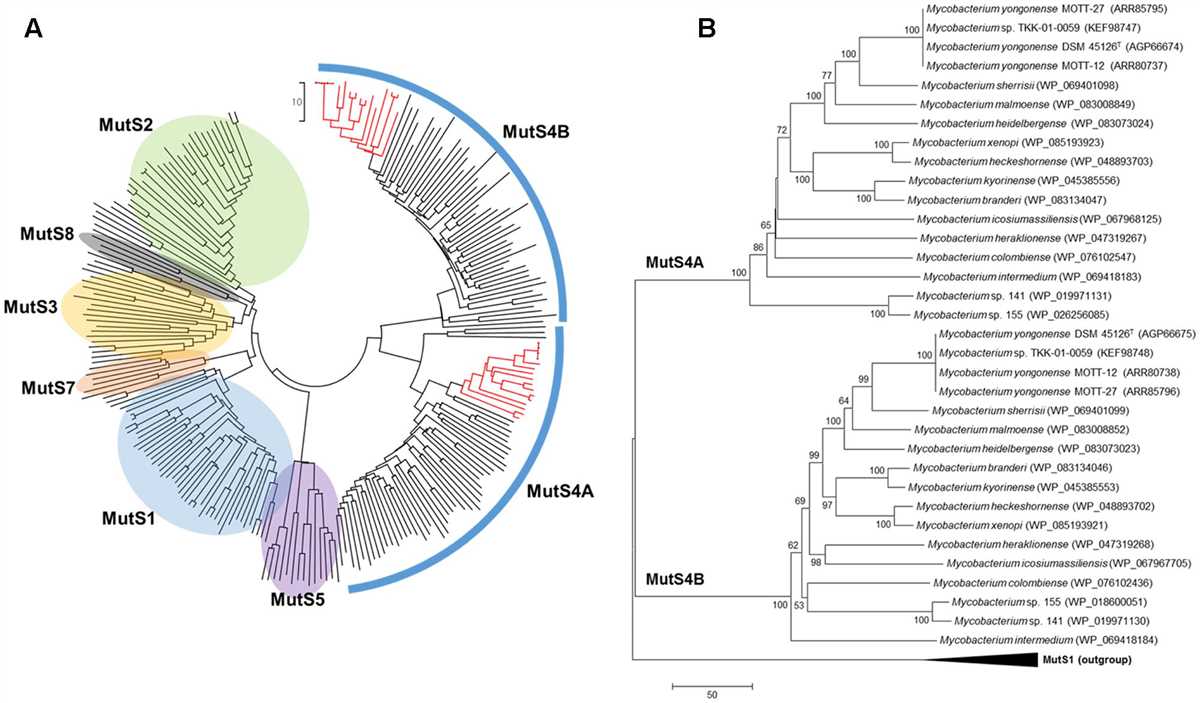
To understand phylogenetic trees, it is important to be familiar with some key concepts and terminology. One of the central concepts is the idea of a common ancestor, which is a hypothetical ancestral organism from which two or more descendant species or groups have evolved. The branches on a phylogenetic tree represent these descendant species or groups, while the nodes represent the common ancestor.
Another important concept is that of evolutionary distance or divergence, which measures the genetic differences between species. This distance is often quantified using methods such as sequence alignment and calculating the number of differences between DNA sequences. The closer two species are on a phylogenetic tree, the more genetically similar they are.
Building Phylogenetic Trees
Constructing phylogenetic trees involves several steps. The first step is to gather genetic data, such as DNA sequences, from the species of interest. These sequences are then aligned and compared to identify similarities and differences. Based on this information, a matrix of distances or differences between the sequences is created.
Next, various methods, such as neighbor-joining or maximum likelihood, are used to analyze the distance matrix and construct a tree that minimizes the total amount of evolutionary change required. These methods use algorithms to calculate the most likely evolutionary relationships among the species based on the genetic data.
The resulting phylogenetic tree is a graphical representation of the evolutionary history of the species, with the branches indicating the relationships and the lengths of the branches representing the evolutionary distances between the species. This tree can then be used to study patterns of evolution, make predictions about the characteristics of common ancestors, and infer the evolutionary processes that have shaped the diversity of life on Earth.
What are phylogenetic trees and why are they important?
Phylogenetic trees are diagrams that represent the evolutionary relationships between different species. They are important tools in understanding the evolutionary history and biodiversity of life on Earth. By analyzing the similarities and differences in DNA sequences or other genetic markers, scientists can construct phylogenetic trees to determine the ancestral relationships among species and how they are related to a common ancestor. Phylogenetic trees provide a visual representation of the evolutionary history and can help answer questions about species diversity, evolution, and the classification of organisms.
One of the key reasons why phylogenetic trees are important is that they provide a framework for understanding the evolutionary relationships between species. By studying these trees, scientists can identify patterns of common ancestry and trace the origins of different traits and characteristics. This information is crucial for understanding the diversity of life and how different species have evolved over time.
Phylogenetic trees also have practical applications in fields such as medicine and conservation. Understanding the evolutionary relationships between different species can help in the development of treatments and vaccines for diseases, as well as in the conservation of endangered species. By knowing the evolutionary history of a species, scientists can make informed decisions about its conservation status and prioritize conservation efforts.
In addition, phylogenetic trees can also help in the understanding of the spread and evolution of diseases. By comparing the DNA sequences of different strains of a pathogen, scientists can construct phylogenetic trees to track the transmission and evolution of the disease. This information can be crucial in developing strategies to control and treat infectious diseases.
In summary, phylogenetic trees are important tools in understanding the evolutionary history and relationships between different species. They provide a visual representation of evolution and help answer questions about biodiversity, evolution, and conservation. In fields such as medicine and disease control, phylogenetic trees play a crucial role in understanding the spread and evolution of diseases.
DNA Sequences and Phylogenetic Trees
DNA sequences are the building blocks of life, containing the genetic information that determines the traits and characteristics of organisms. By comparing the DNA sequences of different organisms, scientists can uncover their evolutionary relationships and construct phylogenetic trees.
Phylogenetic trees are diagrams that depict the evolutionary relationships between different species or groups of organisms. These trees are created based on the similarities and differences in their DNA sequences. Each branch of the tree represents a common ancestor, and the length of the branches indicates the amount of genetic divergence between species.
To create a phylogenetic tree from DNA sequences, scientists first align the sequences of the genes or regions of interest. This alignment allows them to identify the similarities and differences in the sequences. Once the sequences are aligned, scientists can use various algorithms and statistical methods to determine the most likely evolutionary relationships between the organisms.
Creating a phylogenetic tree is a complex process that requires careful analysis and interpretation of the DNA sequences. It requires knowledge of genetics, bioinformatics, and evolutionary biology. By studying the DNA sequences and constructing phylogenetic trees, scientists gain insights into the evolutionary history of organisms and can make predictions about their shared ancestry and evolutionary patterns.
- DNA sequences are the building blocks of life
- Phylogenetic trees are diagrams that depict the evolutionary relationships between different species
- Alignment of DNA sequences is crucial in creating phylogenetic trees
- Scientists use algorithms and statistical methods to determine evolutionary relationships
How are DNA sequences used to create phylogenetic trees?
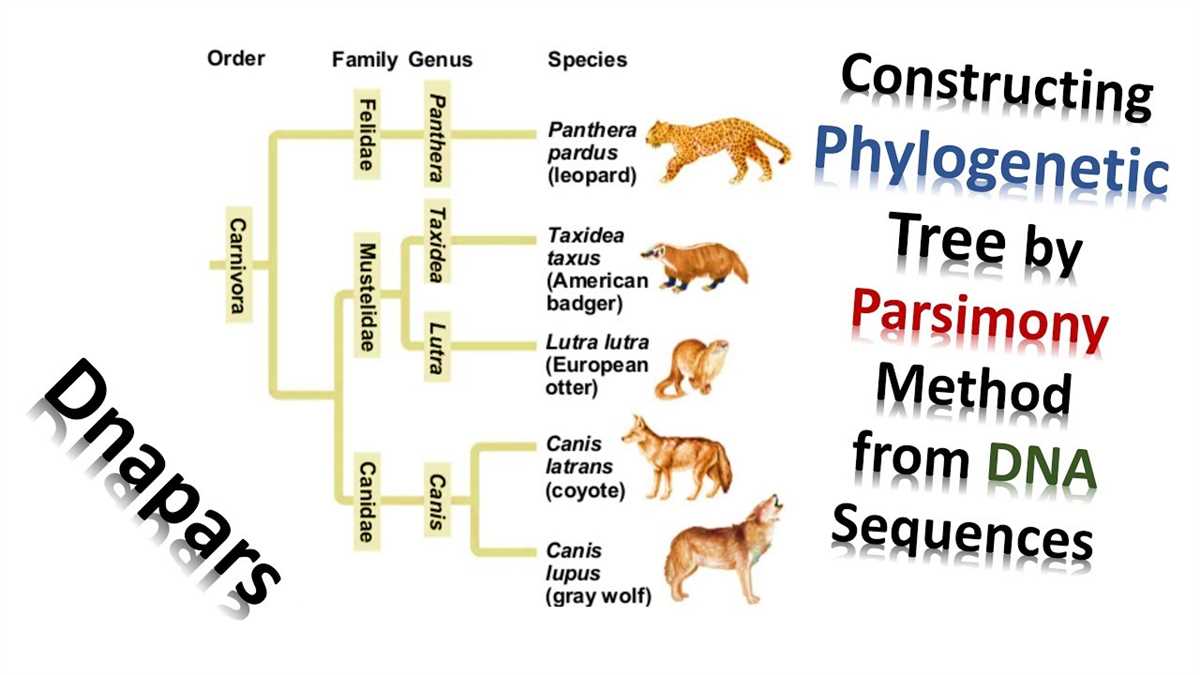
DNA sequences are an essential tool in creating phylogenetic trees, which are diagrams used to show the evolutionary relationships between different species. By comparing DNA sequences from different organisms, scientists can determine how closely related they are and construct a tree that represents their evolutionary history.
One common method used to create phylogenetic trees is called sequence alignment. This involves aligning the DNA sequences of different species and identifying similarities and differences. By analyzing these similarities and differences, scientists can infer how closely related the species are and how they have evolved over time.
Another important technique is molecular clock analysis. This method uses DNA sequences to estimate how long ago species diverged from a common ancestor. By comparing the number of differences in their DNA sequences, scientists can calculate the rate at which mutations occur and use this information to determine the time of divergence.
To construct a phylogenetic tree, scientists analyze the DNA sequences of multiple genes or specific regions of the genome. They use various computational methods, such as maximum likelihood or Bayesian inference, to infer the most likely evolutionary relationships between the species. These methods take into account the patterns of similarities and differences in the DNA sequences and generate a tree that represents the best-fit evolutionary history.
In summary, DNA sequences provide valuable information for creating phylogenetic trees. They allow scientists to analyze the similarities and differences between species, estimate their evolutionary divergence, and infer their evolutionary relationships. By combining DNA analysis with computational methods, scientists can construct phylogenetic trees that provide insights into the evolutionary history of different organisms.
Worksheet Answers for Creating Phylogenetic Trees
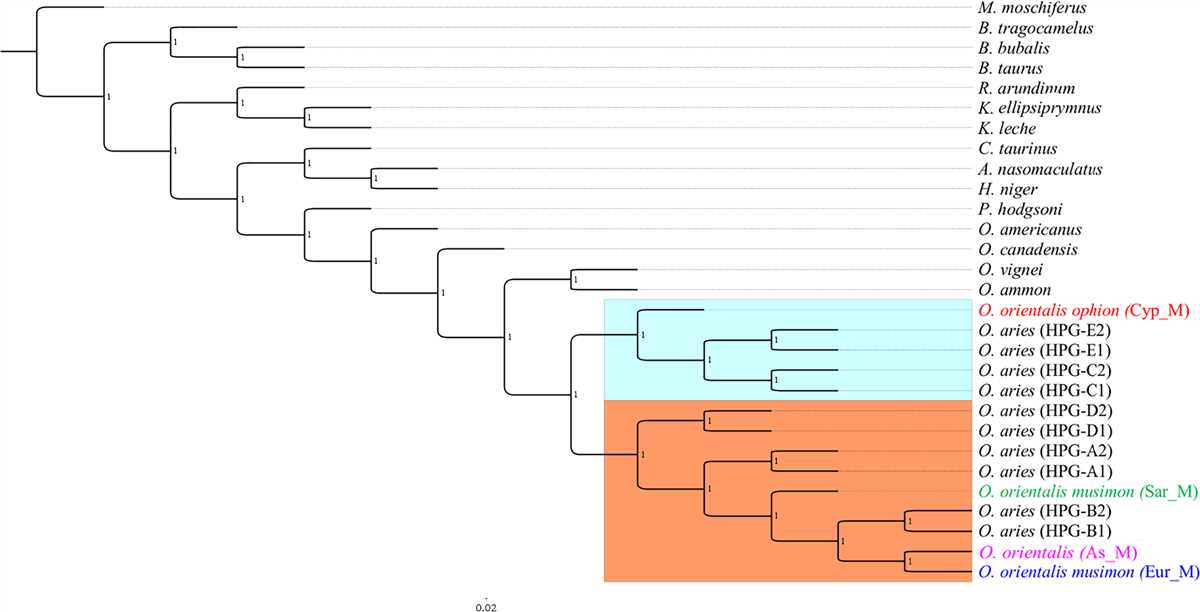
Creating phylogenetic trees from DNA sequences involves analyzing the genetic similarities and differences among different organisms to determine their evolutionary relationships. By comparing the DNA sequences of different species, scientists can construct a tree-like diagram that represents the evolutionary history and common ancestors of these organisms.
To create a phylogenetic tree, scientists first obtain DNA sequences from the organisms they are studying. This could involve extracting DNA from a variety of sources, such as blood samples, tissue samples, or even ancient DNA from preserved specimens. Once the DNA sequences are obtained, they are aligned to identify the corresponding positions of nucleotides in each sequence.
- Question 1: What is the purpose of creating phylogenetic trees from DNA sequences?
- Question 2: How do scientists obtain DNA sequences for constructing phylogenetic trees?
- Question 3: What is the process of creating a phylogenetic tree?
- Question 4: How can creating phylogenetic trees help in understanding the evolution of organisms?
The purpose of creating phylogenetic trees from DNA sequences is to determine the evolutionary relationships between different organisms and understand their shared ancestry.
Scientists obtain DNA sequences by extracting DNA from various sources, such as blood samples, tissue samples, or preserved specimens. The DNA is then sequenced to determine the nucleotide sequence.
The process of creating a phylogenetic tree involves aligning DNA sequences from different organisms and determining their similarities and differences. This information is used to construct a tree diagram that represents the evolutionary relationships and common ancestry of these organisms.
Creating phylogenetic trees helps in understanding the evolution of organisms by providing insights into how different species are related and how they have evolved over time. It allows scientists to trace the evolutionary history and common genetic ancestors of organisms.
Step-by-step guide to completing the worksheet
The worksheet on creating phylogenetic trees from DNA sequences is designed to help you understand the process of using genetic data to construct evolutionary trees. By following these step-by-step instructions, you will be able to successfully complete the worksheet and gain a deeper understanding of phylogenetics.
Step 1: Read the instructions
Begin by carefully reading the instructions provided on the worksheet. Make sure you understand the overall objectives and the specific tasks you need to complete.
Step 2: Obtain the DNA sequences
The worksheet will provide you with a set of DNA sequences as well as information about the species they belong to. Take note of this information as it will be crucial for constructing the phylogenetic tree.
Step 3: Align the DNA sequences
For the next step, you will need to align the DNA sequences using a bioinformatics tool or software. Ensure that the sequences are aligned properly to account for any gaps or mismatches that may occur.
Step 4: Apply a model of evolution
Select a suitable model of DNA evolution to apply to the aligned sequences. This model will describe the different mutation rates and patterns that occur within the sequences. Choose a model that best fits your data and the known characteristics of the DNA sequences.
Step 5: Construct a phylogenetic tree
Using the aligned and modeled DNA sequences, you can now begin constructing a phylogenetic tree. There are various methods and algorithms available for this purpose, such as maximum likelihood or Bayesian methods. Choose a method that you are comfortable with and that suits your data.
Step 6: Analyze and interpret the results
Once you have constructed the phylogenetic tree, analyze and interpret the results. Pay attention to the tree’s branching patterns and the relationships between the species. Consider the evolutionary distances between the sequences and how they relate to the overall tree topology.
By following these six steps, you will have successfully completed the worksheet on creating phylogenetic trees from DNA sequences. This exercise will help enhance your understanding of how genetic data can be used to infer evolutionary relationships and uncover the history of life on Earth.
Understanding the Role of Alignment in Phylogenetic Analysis
Alignment plays a crucial role in phylogenetic analysis, as it allows researchers to compare DNA sequences across different species and construct evolutionary relationships. By aligning sequences, researchers can identify similarities and differences in genetic material, which provides insights into the evolutionary history of organisms.
Alignment involves arranging DNA sequences so that corresponding positions are aligned, allowing for direct comparison of nucleotides. This process ensures that regions of similarity and variation are accurately represented. In phylogenetic analysis, multiple sequence alignment is commonly used, where sequences from different species are aligned together, allowing for comparative analysis.
Alignment algorithms are used to align sequences, and these algorithms employ various techniques to optimize the alignment based on different criteria. Some algorithms focus on minimizing the number of gaps introduced, while others prioritize preserving conserved regions. These algorithms take into account the potential for mutations, insertions, and deletions, which can occur throughout evolution, and aim to create the most accurate alignment possible.
Accurate alignment is essential for constructing phylogenetic trees as it ensures that the evolutionary relationships inferred from the sequences are reliable. A well-aligned sequence allows researchers to accurately identify homologous regions and make meaningful comparisons. Without proper alignment, incorrect relationships may be inferred, leading to misleading conclusions about evolutionary connections.
In conclusion, alignment plays a crucial role in phylogenetic analysis by allowing for the comparison of DNA sequences and the construction of accurate evolutionary trees. Through alignment, researchers can identify similarities and differences in genetic material, infer evolutionary relationships, and gain insights into the history of life on our planet.
Why is sequence alignment important in creating accurate phylogenetic trees?
Sequence alignment is a crucial step in creating accurate phylogenetic trees as it allows scientists to compare and analyze the DNA sequences of different organisms. By aligning the sequences, scientists can identify similarities and differences in the genetic code, which in turn helps them determine the evolutionary relationships between species.
1. Identification of homologous regions: Sequence alignment helps in identifying homologous regions, i.e., regions of DNA that are derived from a common ancestor. These regions are important for understanding the evolutionary history and relationships between organisms. By aligning sequences, scientists can accurately identify and compare these homologous regions.
2. Inferring evolutionary relationships: Sequence alignment is essential for inferring the evolutionary relationships between organisms. By aligning DNA sequences, scientists can identify shared traits and variations, allowing them to reconstruct the common ancestor and the evolutionary paths of different species. This information is crucial for building accurate phylogenetic trees.
3. Accurate assessment of genetic changes: Sequence alignment helps in accurately assessing the genetic changes that have occurred over time. By aligning sequences, scientists can identify insertions, deletions, and substitutions, which are important indicators of evolutionary changes. This information is necessary for constructing a comprehensive and accurate phylogenetic tree.
Overall, sequence alignment plays a vital role in creating accurate phylogenetic trees by identifying homologous regions, inferring evolutionary relationships, and assessing genetic changes. It allows scientists to compare and analyze DNA sequences, providing insights into the evolutionary history and relationships between different organisms.